For a recent project I needed to calculate the pairwise distances of a set of observations to a set of cluster centers. In MATLAB you can use the pdist function for this. As far as I know, there is no equivalent in the R standard packages. So I looked into writing a fast implementation for R. Turns out that vectorizing makes it about 40x faster. Using Rcpp is another 5-6x faster, ending up with a 225x speed-up over the naive implementation.
Archives
12
May 13
Using C libraries in R with rdyncall
One reason I like using R for data analysis is that R has a great collection of packages that let you easily apply state-of-the-art methods to your problems. But once in a while you find a library that you would like to use that does not have a R wrapper, yet. While the great Rcpp package provides a convenient way to write R extensions in C++, it obviously requires you to write C++ code and to have a compiler installed.
An alternative I found about only recently is the rdyncall package. rdyncall provides an improved Foreign Function Interface (FFI), which allows you to dynamically invoke C libraries.
In this blog post I want to give you an example on how to employ rdyncall to use
the LWPR libary for Locally Weighted Projection Regression.
Continue reading →
06
Nov 12
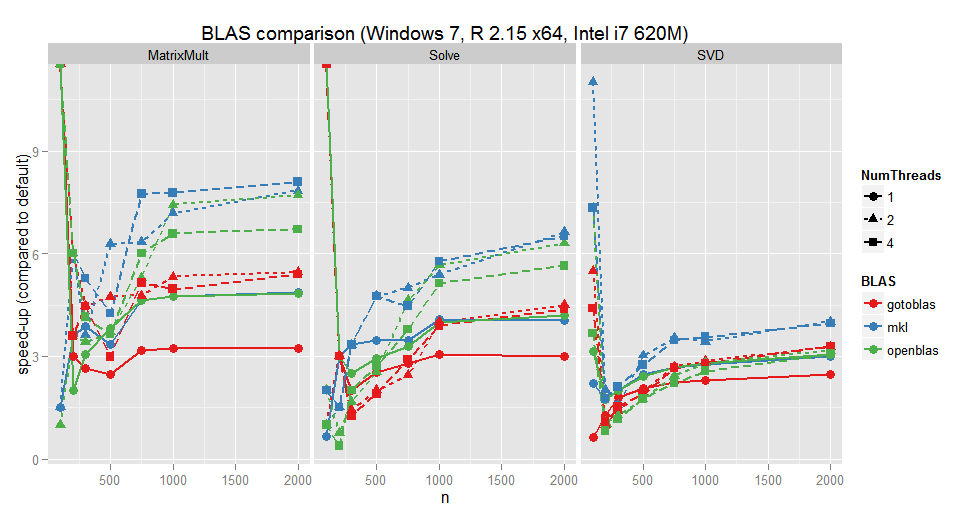
R BLAS: GotoBLAS2 vs OpenBLAS vs MKL
Short update to Speed up R by using a different BLAS implementation/:
- MKL is overall the fastest
- OpenBLAS is faster than its parent GotoBLAS and comes close to MKL
A = matrix(rnorm(n*n),n,n) A %*% A solve(A) svd(A)