One reason I like using R for data analysis is that R has a great collection of packages that let you easily apply state-of-the-art methods to your problems. But once in a while you find a library that you would like to use that does not have a R wrapper, yet. While the great Rcpp package provides a convenient way to write R extensions in C++, it obviously requires you to write C++ code and to have a compiler installed.
An alternative I found about only recently is the rdyncall package. rdyncall provides an improved Foreign Function Interface (FFI), which allows you to dynamically invoke C libraries.
In this blog post I want to give you an example on how to employ rdyncall to use
the LWPR libary for Locally Weighted Projection Regression.
Function signatures
The minimal set of functions we need from the LWPR library consists of lwpr_init_model() to initialize a new model, lwpr_update() to train the model and lwpr_predict() to get predictions from the model.
In the lwpr.h header file you will find the following definitions for these functions:
int lwpr_init_model(LWPR_Model *model, int nIn, int nOut,
const char *name);
int lwpr_update(LWPR_Model *model, const double *x,
const double *y, double *yp, double *max_w);
void lwpr_predict(const LWPR_Model *model, const double *x,
double cutoff, double *y, double *conf,
double *max_w);
In order to use these functions with rdyncall we have to translate the C function definitions into rdyncall’s function signature syntax.
The syntax is function_name(argument_types)return_type;. Most types are respresented by single letters: i = integers, d = double, Z = const char*, … Pointer types are prepended by an asterisk (*), so *d stands for double* in C. Custom types (structs) can also be used and pointers to structs are written as *. The package documentation provides a complete list of available type notations. For our three functions above the signatures for rdyncall look like this:
lwpr_init_model(*<LWPR_Model>iiZ)i; lwpr_update(*<LWPR_Model>*d*d*d*d)i; lwpr_predict(*<LWPR_Model>*dd*d*d*d)v;
Struct definition
In the function signatures we used the struct LWPR_Model. In order to be able to create and modify LWPR_Model “objects” we also have to specify the memory layout of the C struct. The notation for struct definitions is StructName{field_tyoes}fieldname1 fieldname2 ...;.
The C definition of LWPR_Model is:
typedef struct LWPR_Model {
int nIn; /**< \brief Number N of input dimensions */
int nInStore; /**< \brief Storage-size of any N-vector, for aligment purposes */
int nOut; /**< \brief Number M of output dimensions */
int n_data; /**< \brief Number of training data the model has seen */
double *mean_x; /**< \brief Mean of all training data the model has seen (Nx1) */
double *var_x; /**< \brief Mean of all training data the model has seen (Nx1) */
char *name; /**< \brief An optional description of the model (Mx1) */
int diag_only; /**< \brief Flag that determines whether distance matrices are handled as diagonal-only */
int meta; /**< \brief Flag that determines wheter 2nd order updates to LWPR_ReceptiveField.M are computed */
double meta_rate; /**< \brief Learning rate for 2nd order updates */
double penalty; /**< \brief Penalty factor used within distance metric updates */
double *init_alpha; /**< \brief Initial learning rate for 2nd order distance metric updates (NxN) */
double *norm_in; /**< \brief Input normalisation (Nx1). Adjust this to the expected variation of your data. */
double *norm_out; /**< \brief Output normalisation. Adjust this to the expected variation of your output data. */
double *init_D; /**< \brief Initial distance metric (NxN). This often requires some tuning (NxN) */
double *init_M; /**< \brief Cholesky factorisation of LWPR_Model.init_D (NxN) */
double w_gen; /**< \brief Threshold that determines the minimum activation before a new RF is created. */
double w_prune; /**< \brief Threshold that determines above which (second highest) activation a RF is pruned. */
double init_lambda; /**< \brief Initial forgetting factor */
double final_lambda; /**< \brief Final forgetting factor */
double tau_lambda; /**< \brief This parameter describes the annealing schedule of the forgetting factor */
double init_S2; /**< \brief Initial value for sufficient statistics LWPR_ReceptiveField.SSs2 */
double add_threshold;/**< \brief Threshold that determines when a new PLS regression axis is added */
LWPR_Kernel kernel; /**< \brief Describes which kernel function is used (Gaussian or BiSquare) */
int update_D; /**< \brief Flag that determines whether distance metric updates are performed (default: 1) */
LWPR_SubModel *sub; /**< \brief Array of SubModels, one for each output dimension. */
struct LWPR_Workspace *ws; /**< \brief Array of Workspaces, one for each thread (cf. LWPR_NUM_THREADS) */
double *storage; /**< \brief Pointer to allocated memory. Do not touch. */
double *xn; /**< \brief Used to hold a normalised input vector (Nx1) */
double *yn; /**< \brief Used to hold a normalised output vector (Nx1) */
} LWPR_Model;
For dyncall it has to be (manually) transformed to:
LWPR_Model{iiii*d*d*ciidd*d*d*d*d*ddddddddiipp*d*d*d}nIn nInStore nOut n_data mean_x var_x name diag_only meta meta_rate penalty init_alpha norm_in norm_out init_D init_M w_gen w_prune init_lambda final_lambda tau_lambda init_S2 add_threshold kernel update_D sub ws storage xn yn
For fields with pointers to other custom types, such as LWPR_Workspace, we can safely write p for a generic pointer. The enum LWPR_Kernel is represented by an integer (i).
Putting things together
require("rdyncall")
dynbind(
"lwpr", # library name; rdyncall will look for liblwpr.so or lwpr.dll on the library path
"lwpr_init_model(*<LWPR_Model>iiZ)i;
lwpr_update(*<LWPR_Model>*d*d*d*d)i;
lwpr_predict(*<LWPR_Model>*dd*d*d*d)v;
lwpr_duplicate_mode(*<LWPR_Model>*<LWPR_Model>)i;
lwpr_set_init_alpha(*<LWPR_Model>d)i;
lwpr_set_init_D(*<LWPR_Model>*dd)i;
lwpr_set_init_D_diagonal(*<LWPR_Model>*d)i;
lwpr_set_init_D_spherical(*<LWPR_Model>d)i;
lwpr_write_xml(*<LWPR_Model>Z)i;
lwpr_read_xml(*<LWPR_Model>Z*i)i;
")
# create struct type LWPR_Model
parseStructInfos("LWPR_Model{iiii*d*d*ciidd*d*d*d*d*ddddddddiipp*d*d*d}nIn nInStore nOut n_data mean_x var_x name diag_only meta meta_rate penalty init_alpha norm_in norm_out init_D init_M w_gen w_prune init_lambda final_lambda tau_lambda init_S2 add_threshold kernel update_D sub ws storage xn yn")
This code will dynamically load the LWPR library and will create a function in the global environment for each given function signature. The parseStructInfos invocation will create a variable LWPR_Model that can be used with new.struct() to create new LWPR_Model objects.
These 20 lines of R code are enough to put everything in place. We are now able to use the LWPR library in R :-).
The LWPR tutorial code in R
This article would not be complete without an example that actually uses the LWPR library.
So here is the
LWPR tutorial code in R:
testfunc = function(x) {
10 * sin(7.8*log(1+x)) / (1 + 0.1*x**2)
}
Ntr = 500
Xtr = 10 * runif(Ntr)
Ytr = sapply(Xtr, testfunc) + 0.1 * rnorm(Ntr) * Xtr
model = new.struct(LWPR_Model)
lwpr_init_model(model, 1, 1, "Tutorial")
lwpr_set_init_D(model, 20, 0)
model$update_D = 1
model$diag_only = TRUE
model$penalty = 0.0001
lwpr_set_init_alpha(model, 40)
for (i in 1:20) {
idx = sample(Ntr,Ntr)
for (j in idx) {
yp=0
lwpr_update(model, Xtr[j], Ytr[j], yp, NULL)
}
}
Ntest = 500;
Xtest = seq(0,10,length.out=Ntest)
Ytest = numeric(Ntest)
Conf = numeric(Ntest)
for (k in 1:Ntest) {
yp = 0
conf = 0
lwpr_predict(model, Xtest[k], 0.0, yp, conf, NULL);
Ytest[k] = yp
Conf[k] = conf
}
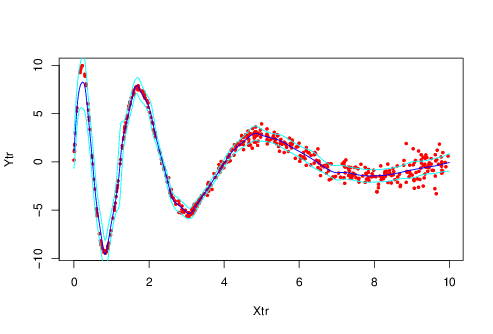
plot(Ytr ~ Xtr, pch=19, cex=0.5, col='red')
lines(Xtest , Ytest, col="blue")
lines(Xtest , Ytest + Conf, col="cyan")
lines(Xtest , Ytest - Conf, col="cyan")
Conclusion
rdyncall allows you to call C library in R without writing any wrapper code. I find it quicker and easier than the alternatives, but it still requires a little knowledge of C to write the function and struct definitions based on the C header files. In the R Journal Article (PDF) about rdyncall a automatic conversion from C header files to rdyncall specifications is mentioned, but I think it was not released.
See Also
- Adler, D.: Foreign Library Interface, R Journal Vol.4/1, June 2012 (PDF)
- Complete example: https://gist.github.com/felixr/5564929
Pretty nice, thanks for the post!